JASPAR Documentation
Last updated: 15 Sep. 2021
JASPAR is a regularly maintained open-access database storing manually curated TF binding preferences as position frequency matrices (PFMs). PFMs summarize nucleotide occurrences at each position in a set of observed TF-DNA interactions. PFMs can be transformed into probabilistic models to construct position weight matrices (PWMs) or position-specific scoring matrices (PSSMs), which then can be used to scan any DNA sequence and predict transcription factors binding sites (TFBSs). The JASPAR database is also providing TFBSs predicted using profiles in the CORE collection.
The motifs in JASPAR are collected in two ways (Figure 1):
- Internally: de novo generated motifs, by analyzing ChIP-seq/-exo sequences using a custom motif discovery pipeline (check the code at our repository).
- Externally: motifs taken directly from other publications and/or resources.
In both cases, the selected motifs are manually curated, which means that our curators found an orthogonal publication giving support to the motif (e.g., a motif found in ChIP-seq peaks looks similar to one found by SELEX-seq). The Pubmed ID associated with the orthogonal support is provided in the TF profile metadata (Figure 1).
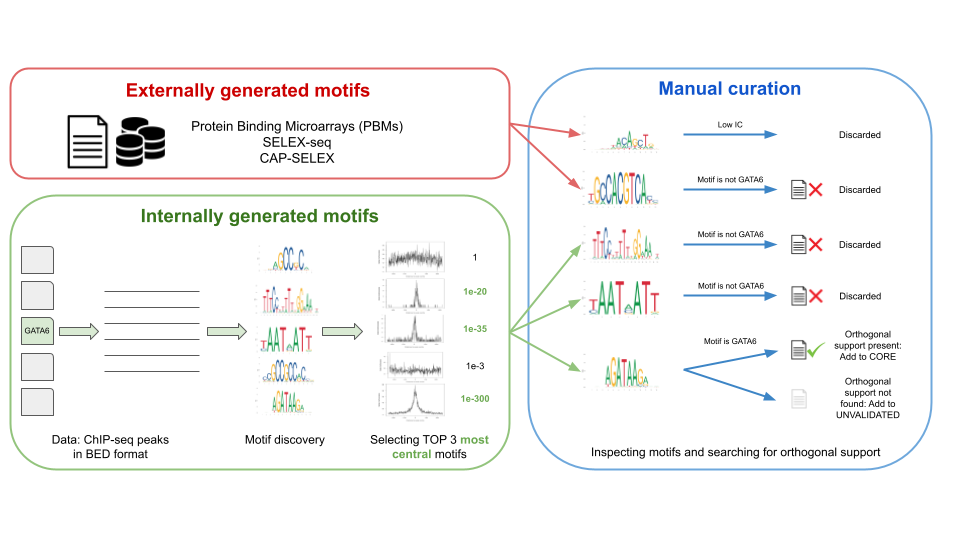
Figure 1. Workflow of data processing and motif curation for the JASPAR database. Motifs in the JASPAR database are of two types: externally and internally generated motifs. Both types of motifs are then passed to the manual curation step, where motifs are manually inspected by a team of curators and orthogonal support in the literature is reviewed.
JASPAR is the only database with this scope where the data can be used without restrictions (open source). For a comprehensive review of models and how they can be used, please see the following reviews:
- Modeling the specificity of protein-DNA interactions Quant Biol. 2013 Jun;1(2):115-130
- Applied bioinformatics for the identification of regulatory elements Nat Rev Genet. 2004 Apr;5(4):276-87
The JASPAR database consists of subsets of profiles known as collections. Each of these collections has different goals, as described below. The main collection is known as the JASPAR CORE and is the collection most scientists use.
JASPAR Collections
The JASPAR database consists of smaller subsets of profiles known as collections. Each of these collections have different goals as described below. The main collection is known as JASPAR CORE and is the collection most scientists use.
JASPAR CORE
The JASPAR CORE collection contains a manually curated, non-redundant set of TF binding profiles. All profiles are derived from published collections of experimentally defined TF binding motifs for eukaryotes. The TF binding profiles were historically determined from SELEX experiments or data collection from the experimentally determined binding regions of actual regulatory regions. More recent profiles are derived from high-throughput techniques such as ChIP-seq/-exo, Protein Binding Microarray, DAP-seq, or High-Throughput SELEX. One of the central goals of the JASPAR CORE is to provide a single “good” model for each TF. This means that the database is non-redundant because there is ideally only one model per TF. There are some exceptions motivated by the recognition of significantly different motifs, where the different profiles are considered as variants. See, for example, CTCF motifs in JASPAR vertebrates for 3 different binding variants.
The JASPAR CORE is what most scientists mean when referring to JASPAR in their manuscripts. For convenience, the JASPAR CORE is divided into taxonomic groups. At the moment, there are profiles for six taxonomic groups: vertebrates, plants, fungi, insects, urochordata, and nematodes. This distinction is mainly used in the web interface and in the download section.
When should it be used? This is the main JASPAR collection and should be used when curated, non-redundant binding profile models for specific factors derived from experimental data are required. TF binding motifs may be used to predict TFBSs on DNA sequences, to investigate the binding preferences of TFs to DNA sequences, or as a reference when annotating/comparing de novo motifs.
Note about the distinction between the taxonomic classification of JASPAR TF binding profiles and the species metadata information:
JASPAR provides TF binding profiles by taxon since TFs with similar DNA binding domains generally share similar binding preferences. The species metadata attached to these profiles indicates the origin of the data used to construct the profiles rather than limiting their applicability to the same species. Therefore, when predicting TF binding sites or studying TF binding preferences, users should consider utilizing all TF binding profiles from the relevant taxon rather than confining themselves to profiles derived solely from the species of interest.
JASPAR UNVALIDATED
The JASPAR UNVALIDATED collection was introduced in the 2020 release of the database. These profiles can be browsed separately on the JASPAR website and can be distinguished by an identifier that starts with “UN” (Figure 2A,B). These profiles are regarded as unvalidated because our curators could not find any orthogonal support from the existing literature. However, the profiles in this collection are high-quality, such as a profile with high IC and good centrality of the associated sites around the ChIP-seq peak summits (see Figure 2C).
We encourage the community to engage in the curation of these profiles by performing experiments and/or pointing us to literature that our curators missed to support these profiles. Suggestions for validation can be submitted via the form provided on the profiles page (see Figure 2D).
When should it be used? These profiles are not non-validated, so we recommend using them with caution.

Figure 2. Screengrab of the DPF1 profile (UN0116.1). The highlighted sections indicate where to browse the UNVALIDATED collection (A) in which the profiles have identifiers always starting with “UN” (B). These profiles do not yet have orthogonal support but are of high quality, e.g. high IC and good centrality (C). The scientific community is invited to participate in the curation of the UNVALIDATED collection by submitting their suggestions for validation (D).
JASPAR Metadata
Each profile when displayed in the website provides all kinds of information. First of all, the metadata describing the profile (see Table 1 and Figure 3). Then, the matrix and motif logo are displayed. At the bottom of the page there is also information available on all versions of the profiles (see Figure 3).
| # | Entry | Note |
|---|---|---|
| 1 | Name | The name of the TF. As far as possible, the name is based on the standardized Entrez gene symbols. In the case the model describes a TF hetero-dimer, two names are concatenated, such as Pou5f1::Sox2. In a few cases, one particular TF may have different splice forms resulting in different binding specificity, in previous releases these cases were handled as binding variants (e.g., CEBPG (MA0838.1) and CEBPG(var.2) (MA1636.1)). From the JASPAR 2022 release, binding variants are not anymore indicated as part of the TF name: all binding variants for the same TF share the TF name, but each one has a different Matrix ID. |
| 2 | Matrix ID | A unique identifier for each model. CORE matrices always have identifiers starting with “MA”, while profiles in the UNVALIDATED collection have identifiers starting with “UN”. The number after the dot (e.g., MA1636.1) corresponds to the version. Larger number indicates that a motif has been updated. |
| 3 | Class | Structural class of the transcription factor, based on the TFClass system. |
| 4 | Family | Structural sub-class of the transcription factor, based on the TFClass system. |
| 5 | Collection | Indicating which collection the profile belongs: CORE or UNVALIDATED. |
| 6 | Taxon | Group of species, currently consisting of 6 larger groups: vertebrate, plants, fungi, insects, urochordata and nematodes. |
| 7 | Species | The species source for the sequences, in Latin. Linked to the NCBI Taxonomic browser. The actual database entries are the NCBI tax IDs – the latin conversion is only in the web interface. |
| 8 | Data type | Methodology used for matrix construction, e.g., ChIP-seq, PBM, SELEX. |
| 9 | Validation | A link to PubMed indicating the orthogonal evidence of the Tf binding profile. |
| 10 | Uniprot ID | A link to the corresponding UniProt record. |
| 11 | Source | A reference to the data, which was used to build a profile and where the profile was taken from. |
| 12 | Comment | For some matrices, a curator comment is added. |
Table 1. The metadata categories that are displayed in a profile page. Each category of the metadata has a number assigned and is also shown in Figure 3.

Figure 3. GATA2 profile in JASPAR (MA0036.3), highlighting the metadata and other information available. Metadata categories (green numbers) are explained in more detail in Table 1.
JASPAR Matrix formats
JASPAR stores transcription factor binding profiles in four formats. Following is more information on formats and the DNA binding profile for GATA6 transcription factor (JASPAR ID MA1104.2) as an example:
Raw PFM
Each matrix is separated by a FASTA-like header starting with the > symbol and then a matrix ID. The count for each base (ACGT) is specified on its own space separated line where each element corresponds to one column. The order of the lines for the bases is A, C, G and finally T.
>MA1104.2 GATA6
22320 20858 35360 5912 4535 2560 5044 76686 1507 1096 13149 18911 22172
16229 14161 13347 11831 62936 1439 1393 815 852 75930 3228 19054 17969
13432 11894 10394 7066 6459 580 615 819 456 712 1810 18153 11605
27463 32531 20343 54635 5514 74865 72392 1124 76629 1706 61257 23326 27698
JASPAR
This is similar to the raw format, having an identical header. The lines for each base however start with a label for the nucleotide (A, C, G or T) and then the columns follow enclosed in brackets: [].
>MA1104.2 GATA6
A [ 22320 20858 35360 5912 4535 2560 5044 76686 1507 1096 13149 18911 22172 ]
C [ 16229 14161 13347 11831 62936 1439 1393 815 852 75930 3228 19054 17969 ]
G [ 13432 11894 10394 7066 6459 580 615 819 456 712 1810 18153 11605 ]
T [ 27463 32531 20343 54635 5514 74865 72392 1124 76629 1706 61257 23326 27698 ]TRANSFAC
This is a TRANSFAC-like format having a few lines with information, such as “AC”, which stores JASPAR matrix unique ID, “ID” indicates the TF name and “DE” has both. The data itself is transposed as compared to the other formats, meaning that each line corresponds to a column in the profile. The column lines start with a number denoting the column index (counting from 0). Additional lines starting with “CC” store some additional metadata, such as TF family and class. A final line of the matrix file is indicated with “//”. Empty lines are indicated with “XX”.
AC MA1104.2
XX
ID GATA6
XX
DE MA1104.2 GATA6 ; From JASPAR
PO A C G T
01 22320.0 16229.0 13432.0 27463.0
02 20858.0 14161.0 11894.0 32531.0
03 35360.0 13347.0 10394.0 20343.0
04 5912.0 11831.0 7066.0 54635.0
05 4535.0 62936.0 6459.0 5514.0
06 2560.0 1439.0 580.0 74865.0
07 5044.0 1393.0 615.0 72392.0
08 76686.0 815.0 819.0 1124.0
09 1507.0 852.0 456.0 76629.0
10 1096.0 75930.0 712.0 1706.0
11 13149.0 3228.0 1810.0 61257.0
12 18911.0 19054.0 18153.0 23326.0
13 22172.0 17969.0 11605.0 27698.0
XX
CC tax_group:vertebrates
CC tf_family:GATA-type zinc fingers
CC tf_class:Other C4 zinc finger-type factors
CC pubmed_ids:9915795
CC uniprot_ids:Q92908
CC data_type:ChIP-seq
XX
//MEME
MEME motif format is a simple text format for motifs that is accepted by the programs in the MEME Suite that require MEME Motif Format. A text file in MEME minimal motif format can contain more than one motif, and also (optionally) specifies the motif alphabet, background frequencies of the letters in the alphabet, and strand information (for motifs of complementable alphabets like DNA), as illustrated in the example below:
MEME version 4
ALPHABET= ACGT
strands: + -
Background letter frequencies
A 0.25 C 0.25 G 0.25 T 0.25
MOTIF MA1104.2 GATA6
letter-probability matrix: alength= 4 w= 13 nsites= 79444 E= 0
0.280953 0.204282 0.169075 0.345690
0.262550 0.178251 0.149716 0.409483
0.445093 0.168005 0.130834 0.256067
0.074417 0.148923 0.088943 0.687717
0.057084 0.792206 0.081303 0.069407
0.032224 0.018113 0.007301 0.942362
0.063491 0.017534 0.007741 0.911233
0.965284 0.010259 0.010309 0.014148
0.018969 0.010725 0.005740 0.964566
0.013796 0.955768 0.008962 0.021474
0.165513 0.040632 0.022783 0.771071
0.238042 0.239842 0.228501 0.293616
0.279090 0.226184 0.146078 0.348648
URL https://jaspar.uio.no/matrix/MA1104.2
Version control
Since the 4th release of JASPAR, all matrix models have versions. This is primarily to keep track of improvements - which can be anything from correcting typos to actually making a new model based on new data. Version control works as follows: IDs are based on a stable ID, and a version number, so that the whole ID is [stable ID].[version]. The stable ID follows a certain TF, or other logic unit such as a dimer pair. For instance, the stable ID for the factor GATA2 is MA0036.3. However, the GATA2 matrix has been updated two times with new data, so there are currently four versions: MA0036.1, MA0036.2, and MA0036.3 (See Figure 3).
The latest versions of all profiles with orthogonal support are referred to as JASPAR CORE non-redundant collection (latest versions of profiles without orthogonal support are stored in JASPAR UNVALIDATED non-redundant). All versions of profiles are also available in the database and referred to as redundant collection (CORE and UNVALIDATED).
Accessing and Downloading JASPAR Data
Depending on what the user wants to download, there are several ways to access and download the JASPAR data. TF binding profiles can be downloaded in bulk from the website (all together or per taxon), and metadata can be retrieved using the JASPAR RESTful API or accessed through pyJASPAR. Profiles can also be accessed through the R/Bioconductor package.
JASPAR downloads
The JASPAR collections in bulk are generally downloadable in three different forms from the JASPAR downloads page (Figure 4):
- Flat files in JASPAR, MEME, and TRANSFAC formats for the CORE redundant and non-redundant collections of matrix profiles. The CORE collection is available to download for each specific taxonomic group (Figure 4B,H).
- Archives of files (also in JASPAR, MEME, and TRANSFAC formats for the CORE and UNVALIDATED collections), each providing individual matrix profile information (Figure 4D).
- A database/SQL dump of the data and metadata (Figure 4F).

Figure 4. Screengrab of JASPAR Download page. The download page is accessible through the left panel (A). By default "CORE PFMs" download page is displayed (B), where a user can find all CORE PFMs or for each of the taxonomic groups for download as individual files or a single file (H). The UNVALIDATED collection can be downloaded from JASPAR collections (PFMs) (D). Other data, such as TFFMs, matrix clustering or the entire SQL dump, can also be downloaded (C, E, F). TFBSs and matrix clustering are available under "Other data" (G).
Other sources and analysis results are also available in data files:
- Transcription Factor Flexible Models (TFFMs) are available to download as tar files. Additional information regarding TFFMs is available as a .csv file (Figure 4C).
- Matrix clustering of CORE and CORE + UNVALIDATED results are available to download as radial trees or as clustering summaries for each taxonomic group (Figure 4G).
- Other data, such as sequences used to generate PFMs in FASTA format, genomic coordinates of the sequences in BED format, motif logos and centrality plots, are available as a bulk download under "Other data" (Figure 4G).
JASPAR RESTful API
Since 2020, JASPAR has a Representational State Transfer (REST) application programming interface (API) to query/retrieve matrix profile data from the JASPAR database. It is a browsable API and comes with a human browsable interface and also a programmatic interface, which returns the results in JSON format. For more details, please read the API documentation. If you wish to cite the JASPAR REST API, please check the FAQ page.
pyJASPAR
pyJASPAR is a Pythonic interface to access JASPAR transcription factor profiles. It uses Biopython and SQLite3 to provide a serverless interface to the JASPAR database to query and access TF motifs across current and previous database releases. Currently, the releases available in pyJASPAR are: JASPAR2014, JASPAR2016, JASPAR2018, JASPAR2020, JASPAR2022, and JASPAR2024. Information on the installation of pyJASPAR can be found under Tools , in the pyJASPAR repository and documentation.
R Bioconductor Package
Different releases of JASPAR can also be accessed through Bioconductor data packages. Currently five JASPAR releases are available:
To browse the database TFBSTools package is required. You can find more information about this package and installation under Tools and here.
Browsing JASPAR motif clusters
Since JASPAR 2018, we have introduced a visual representation of all motifs (CORE collection and CORE + UNVALIDATED collections) as a hierarchical tree displaying a global motif alignment (Figure 5). Clusters are accessible by clicking on the “Matrix Clusters” button at the left menu on the JASPAR website (Figure 5A). For each taxon, separately, all motifs are clustered and aligned using the RSAT matrix-clustering tool; these radial trees can be zoomed and explored more closely. For the current release (JASPAR2024), we relied on a standalone version of the RSAT matrix-clustering tool to speed up the clustering task. Details about the standalone RSAT matrix-clustering tool can be found in this repository.

Figure 5. Browsing matrix clustering in JASPAR. All motifs corresponding to a taxonomic group are displayed as a hierarchical tree with a global alignment. A) The TF class information is displayed as an outer ring. B) Users can change motif logo orientation and trees can be zoomed in to ease the motif exploration. C) Each TF name in the tree has a link pointing to its corresponding profile page on the JASPAR website.
The TF classes obtained from TFclass are displayed in the annotation table, each one with an associated number and a colour (Figure 5A). The tree is surrounded by an outer ring containing the TF structural class colours and numbers (Figure 5B). The UNVALIDATED motifs are depicted with a star (*) next to their TF class (Figure 5B). Users can change the TF logo alignment orientation by clicking on the Direct/Reverse red buttons on top (Figure 5B).
The branch colours correspond to the clusters found by the matrix clustering tool, note that although TFs from the same family tend to be grouped in the same cluster, sometimes members of other TF families are grouped together (Figure 5B, see cluster containing TF from classes 9 and 10). The tree leaves correspond to TF names, by clicking on one of them, it will open a tab in your browser with the dedicated profile page (Figure 5D).
JASPAR Software Tools
JASPAR contains not only collections of TF binding profiles but also a suite of tools, such as the TFBS enrichment tool. For further details please check the Tools page.
JASPAR TFBS extraction tool
The JASPAR 2024 release comes with a new TFBS extraction computation tool to extract predicted JASPAR TFBSs that intersect with a set of genomic regions the user provides. It allows TFBS filtering by TF names, JASPAR matrix IDs, and TFBS score thresholds. This software is available as a command-line tool at URL and in a Docker container at URL.
Profile Inference
The motif inference tool in the JASPAR database predicts the probable transcription factor (TF) binding profile recognized by a given protein. This is achieved by comparing the DNA binding domain (DBD) of the provided protein, either as a full amino acid sequence or specifically as the DBD sequence, with the DBDs of all TFs, catalogued in JASPAR. The tool capitalizes on the principle that TFs sharing similar DBDs usually exhibit akin binding preferences. For further insights, the JASPAR 2016 manuscript provides comprehensive details on this process.
Matrix Align
The matrix align tool aligns DNA binding profiles. It takes custom matrices in JASPAR or PFM formats and aligns them to identify the JASPAR profiles that best match the input. The Matrix alignment can be restricted to (1) the CORE or the UNVALIDATED collections, (2) different taxonomic groups, and (3) the latest non-redundant or all (redundant) versions of the TF-binding profiles.